
PSEUDO is a computational framework for debiasing and experimental uncertainty quantification in protein structural models resolved by molecular replacement.
It provides a three-stage pipeline that runs on HPC clusters via SLURM:
| Stage | Command | What it does |
|---|---|---|
| Debias | pseudo-debias |
Generates stochastic omit perturbation (STOMP) maps via Phenix |
| Quantify | pseudo-quantify |
Separates true signal from phase bias across the map ensemble |
| Analyse | pseudo-analyse |
Scores every model atom against the debiased SNR map (MUSE) |
Installation
git clone https://github.com/hamlet-khachatryan/PSEUDO.git
cd PSEUDO
pip install -e ".[dev]"
PSEUDO requires the Phenix Software Suite for STOMP map calculation. Load it before submitting any SLURM jobs:
module load phenix
Minimal configuration
Create a YAML file (e.g. run.yaml) with just the required fields:
debias:
run_name: "my_experiment"
structure_path: "/data/target.pdb"
reflections_path: "/data/target.mtz"
omit_type: "atoms"
omit_fraction: 0.1
iterations: 5
seed: 42
paths:
work_dir: "/scratch/my_project"
slurm:
partition: "cs05r"
All other parameters fall back to built-in defaults (see Configuration Reference).
Input modes
PSEUDO accepts three input modes for the debias stage:
| Mode | Required fields | Behaviour |
|---|---|---|
| Single structure | structure_path + reflections_path |
Always processed as-is |
| CSV screening | screening_path (.csv) |
Must contain PDB/CIF and MTZ columns (all rows always processed) |
| SQLite screening | screening_path (.sqlite) |
Diamond SoakDB format, supports outcome filtering and structure count capping |
SQLite-specific options
sqlite_outcomes — comma-separated substrings matched against the RefinementOutcome column. Accepted values:
CompChem ready, Deposition ready, Deposited
max_structures — cap on the number of structures processed from the SQLite file. Set to null (default) to process all matching entries.
Example config for SQLite with filtering:
debias:
screening_path: "/data/soakdb.sqlite"
sqlite_outcomes: "CompChem ready, Deposition ready, Deposited"
max_structures: 50
Or via CLI:
pseudo-debias generate-params --config run.yaml \
--sqlite_outcomes "CompChem ready, Deposited" \
--max_structures 50
Basic usage
Three commands take a structure from raw reflections to per-atom density support scores:
pseudo-debias generate-params --config run.yaml
# The exact command is printed by generate-params.
# Small runs (omission jobs ≤ screening_chunk_size):
jid=$(sbatch --parsable submit_preprocessing.slurm)
jid=$(sbatch --parsable --dependency=afterok:$jid submit_omission.slurm)
# Large screening runs:
jid=$(sbatch --parsable submit_preprocessing.slurm)
jid=$(sbatch --parsable --dependency=afterok:$jid submit_omission_0.slurm)
jid=$(sbatch --parsable --dependency=afterok:$jid submit_omission_1.slurm)
# …
pseudo-quantify --input_path /scratch/my_project/my_experiment
pseudo-analyse --input_path /scratch/my_project/my_experiment
Quantification results are stored in <work_dir>/<run_name>/<crystal>/quantify_results/<k_*_cap_*>/:
| File | Description |
|---|---|
{stem}_mean.ccp4 |
STOMP$_{\mu}$ map: average over the STOMP ensemble |
{stem}_std.ccp4 |
STOMP$_{\sigma}$ map: voxel-wise standard deviation map of the STOMP ensemble |
{stem}_snr.ccp4 |
STOMP$_{SNR}$ map: voxel-wise signal-to-noise ratio map |
{stem}_p_values.ccp4 |
voxel-wise signal probability map derived from the STOMP$_{SNR}$ map |
Analysis results land in <work_dir>/<run_name>/<crystal>/analyse_results/:
| File | Description |
|---|---|
{stem}_atoms.csv |
Per-atom MUSE scores and diagnostic flags |
{stem}_residues.csv |
Per-residue MUSEm aggregated scores |
{stem}_summary.json |
Global statistics (OPIA, counts, thresholds) |
{stem}_scored.pdb |
Structure with MUSE scores in the B-factor column |
Load {stem}_scored.pdb in PyMOL and colour by B-factor to visualise density support across the model.
Screen Report
When pseudo-analyse processes a screening run (a directory containing multiple crystals), it automatically generates an interactive HTML summary report:
<screening_dir>/index.html
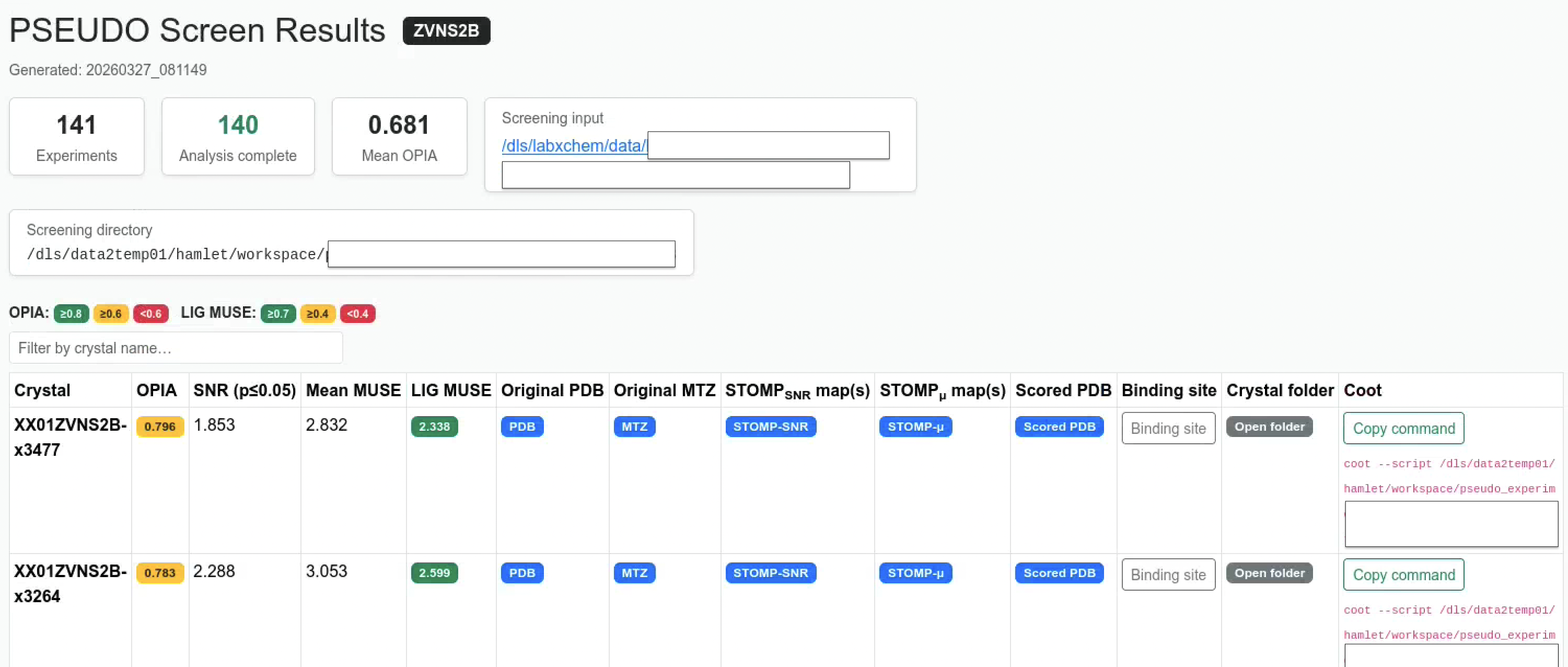
The report collects all crystal results into one table with OPIA and ligand MUSE badges, direct links to maps and structures and pre-built Coot session scripts that load maps for visual inspection.
The report can be regenerated at any time from existing analysis results:
pseudo-screen-report --input_path /scratch/results/my_screen
See the Analyse guide for a full description of every column and the Coot integration.
Logging
Each pipeline stage writes structured eliot logs in NDJSON format to:
<work_dir>/<run_name>/logs/eliot/<name>.ndjson
For example, a debias run named my_experiment produces:
/scratch/my_project/my_experiment/logs/eliot/my_experiment.ndjson
Quantify and analyse stages write one log file per crystal stem (e.g. XTAL-0001.ndjson).
Inspecting logs with eliot-tree
Use the eliot-tree command-line tool to render the nested action tree in a human-readable format:
eliot-tree /scratch/my_project/my_experiment/logs/eliot/my_experiment.ndjson
For a quantify or analyse log:
eliot-tree /scratch/my_project/my_experiment/XTAL-0001/logs/eliot/XTAL-0001.ndjson
Pass eliot-tree --help to see filtering and formatting options.
Further reading
- Debias guide — STOMP map generation, directory layout, batch screening
- Quantify guide — bias separation algorithm
- Analyse guide — MUSE scoring methodology, configuration
- Configuration reference — every parameter for all three stages
Citation
If you use this code, including STOMP maps, the PSEUDO platform, or MUSE scores — in your research, please use the following citation (preprint available soon):
@software{khachatryan2026pseudo,
title = {PSEUDO: Framework for Phase Uncertainty Estimation of Protein Models},
author = {Khachatryan, Hamlet and Wild, Conor and von Delft, Frank},
year = {2026},
url = {https://github.com/hamlet-khachatryan/PSEUDO}
}
MUSE scores
MUSE adapts the EDIA methodology. If you use MUSE scores, please also cite:
@article{meyder2017edia,
title = {Estimating Electron Density Support for Individual Atoms and Molecular Fragments in X-ray Structures},
author = {Meyder, Agnes and Nittinger, Eva and Lange, Gudrun and Klein, Robert and Rarey, Matthias},
journal = {Journal of Chemical Information and Modeling},
volume = {57},
pages = {2437--2447},
year = {2017},
doi = {10.1021/acs.jcim.7b00391}
}
@article{nittinger2015water,
title = {Evidence of Water Molecules --- {A} Statistical Evaluation of Water Molecules Based on Electron Density},
author = {Nittinger, Eva and Meyder, Agnes and Lange, Gudrun and Klein, Robert and Rarey, Matthias},
journal = {Journal of Chemical Information and Modeling},
volume = {55},
pages = {771--783},
year = {2015},
doi = {10.1021/ci500662d}
}
Copyright
Copyright © 2026 Hamlet Khachatryan, Conor Wild, Frank von Delft.
For enquiries, contact hamlet.khachatryan@ndm.ox.ac.uk.